 The same procedure as every year – the influenza season is coming back. It is a recurring threat we all share at the end of each summer. Although not critically dangerous for healthy people, the flu can pose a serious risk to children, the elderly, pregnant women, and immunocompromised patients. According to the World Health Organization (WHO), the flu causes more than half a million deaths every year from respiratory diseases. The flu is caused by RNA viruses, which pass through the air and enter the body through the nose or the mouth. Their ability to mutate and adapt continually makes it harder and harder to defeat them (Kumar et al., 2018).
The same procedure as every year – the influenza season is coming back. It is a recurring threat we all share at the end of each summer. Although not critically dangerous for healthy people, the flu can pose a serious risk to children, the elderly, pregnant women, and immunocompromised patients. According to the World Health Organization (WHO), the flu causes more than half a million deaths every year from respiratory diseases. The flu is caused by RNA viruses, which pass through the air and enter the body through the nose or the mouth. Their ability to mutate and adapt continually makes it harder and harder to defeat them (Kumar et al., 2018).
Of the four different influenza virus strains, influenza A, B, C, and D, the first two are the most common, and they are responsible for most of the infections and deaths during influenza season. Vaccination is the best preventive measure. However, the switch of the viral antigenic epitopes, the so-called antigen drift, makes it necessary to change the vaccine composition every year (Zhou et al., 2018).
A broad spectrum of treatments is available for infected patients, ranging from symptomatic to antiviral treatments. For the latter, using neuraminidase inhibitors is recommended, as they shorten the duration of illness and reduce the risk of complications. These drugs act on viral enzymes to inhibit the release of virus particles from infected cells. The use of another class of antiviral drugs known as aminoadamantanes, which block the M2-channel to inhibit viral uncoating, is no longer recommended, because the virus has largely become resistant to these medications. It is important to emphasize that the development of resistance to antiviral drugs increasingly represents a significant challenge. New strategies for effective treatment are needed.
Influenza A: Researchers focus on better understanding the ever-changing virus

“Influenza viruses constantly rearrange themselves on a genetic level, which results in changes of the proteins on their surface,” explains Dr. Susann Kummer, a post-doctoral scientist with the Department of Infectious Diseases, Virology, at Heidelberg University Hospital. “They can cross species barriers and quickly adapt to new hosts, different environments, and different conditions. Mutations happen very often. Some of these lead to the death of the virus, but others can contribute to the development of resistance mechanisms against antiviral compounds, or make it possible for the virus to pass from animals to humans.”
Kummer is winner of the 2016 Anita and Friedrich-Reutner Prize awarded by the Medical Faculty of the Heidelberg University. She is studying the interactions between the different influenza A virus strains and their host cells (Kummer et al., 2014). Identifying how the critical components of this complex process interact is essential for understanding viral pathogenicity and for designing therapeutic interventions.
New microscopy technologies create super-resolution images
Conventional microscopy does not allow researchers to observe the molecular processes taking place between viruses (~100 nm) and components of the host cells, as objects smaller than 200–300 nm cannot be accurately resolved due to the diffraction properties of light. However, novel super-resolution microscopy techniques have revealed a nanoscale view of viruses’ replication process and cell biology. These approaches use mathematical models or physical parameters to increase resolution beyond the diffraction limit (MacDonald et al., 2015).
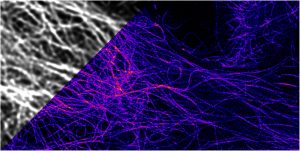
STimulated Emission Depletion (STED) microscopy is the technique used by Kummer. “With STED, having distinct imaging conditions in biological samples means we can achieve a separation sharpness of 20 to 30 nm,” she remarks. “We mark the interesting viral and cell proteins with fluorescent dyes so we can observe their interaction during viral replication. We can also conduct kinetic observations over several minutes.”
This technique was developed by Stefan W. Hell and Jan Wichmann in 1994 and creates super-resolution images by deactivating fluorescence in the surrounding regions and thereby reducing the area of illumination around the focal point. In 2014 Hell, Eric Betzig and W.E. Moerner received the Nobel Prize in Chemistry for their “development of super-resolved fluorescence microscopy.”
In short, STED microscopy is a deterministic method that is based on the scanning of two overlapping light sources, which combine to enable a super-resolved spot on the sample. The stochastic approach developed by Betzig is known as STochastic Optical Reconstruction Microscopy (STORM)), and is based on the precise localization of a large number of individual molecules that form a super-resolved image (Tam and Merino, 2015; Witte et al., 2018). Both techniques allow better insight into the world of proteins, and the choice between STED and STORM depends on the individual research focus.
Recently, Hell and his team at the Max Planck Institute for Biophysical Chemistry in Göttingen have developed a new technique that combines the advantages of STED and STORM, and which allows scientists to discern molecules that are very close to each other with a 1 nm precision (MINFLUX) (Balzarotti et al., 2017). As in STORM, single molecules are randomly switched on and off, and at the same time, their exact position is determined with a “doughnut”-shaped laser beam, as in STED. Images generated with this methodology are more than 100 times sharper than conventional light microscopy, and are up to 20 times sharper than images available with previous super-resolution light microscopy. This breakthrough creates new opportunities for researchers to investigate how viruses behave in human cells. “A lot of small moments of progress during laboratory research suddenly add up to become milestones on the way to bigger scientific discoveries,” comments Kummer.
Primary cells for virology research
In addition to using STED microscopy to inspect the mechanics of viral infection of lung cells, Kummer mimics native conditions using human lung cancer cell lines, as well as primary human small airway epithelial cells (HSAEpC), from PromoCell for her research. “Compared to lung cancer cell lines, primary lung cells allow us to examine different cell types at the same time in a physiological environment. You can see which cell types are preferentially affected by influenza A,” she explains.
To gain even better insights in the pathophysiological processes underlying a viral infection, ongoing research uses three-dimensional structures. “Lung-like organoids will help us to study the distribution of the virus in a three-dimensional structure,” Kummer states. In fact, recent investigations exploring the interaction between avian influenza virus and the intestinal mucosa have shown how viruses can invade intestinal organoids and cause cellular damage (Huang et al., 2017). These studies indicate that three-dimensional structures will be a useful tool for infection disease modeling, in particular as researchers continue to search for ways to defeat influenza viruses.



