In a complicated puzzle, just one piece will fit at each specific place. This is similar to what happens when the cells of our immune system, the T cells, meet an antigen. If they recognize it as a “fitting”, normal antigen, nothing will happen. However, if this is a foreign, or pathologic antigen, T cells will be activated. To be recognized by T cells, antigens must be loaded on human leucocyte antigen (HLA) molecules. HLA is the human version of the major histocompatibility complex (MHC) of vertebrates, a group of cell-surface proteins coded by more than 250 genes on chromosome 6. These are the essential puzzle pieces of the immune-response mechanisms.
All of us inherit many different HLA genes from our parents. HLA types aren’t evenly distributed among the population, as some HLA types are more frequent than others. Our individual HLA type is for example what will reject tissue following organ or stem cell transplantation. “If the HLA type of the donor doesn’t match the HLA type of the recipient, the recipient’s immune system will recognize the transplanted organ, or the transplanted stem cells, as foreign and reject them,” explains Lisa Schmidtke, Scientific Support Specialist at PromoCell. “HLA are also pivotal in cancer immunotherapy, as cancer-specific antigens are presented on HLA molecules. When T cells recognize the antigens of cancer cells, they attack and kill them.” This is why researchers who perform cell culture experiments should know the HLA type of the cells, or risk discovering that the T cells have eliminated the intruders.
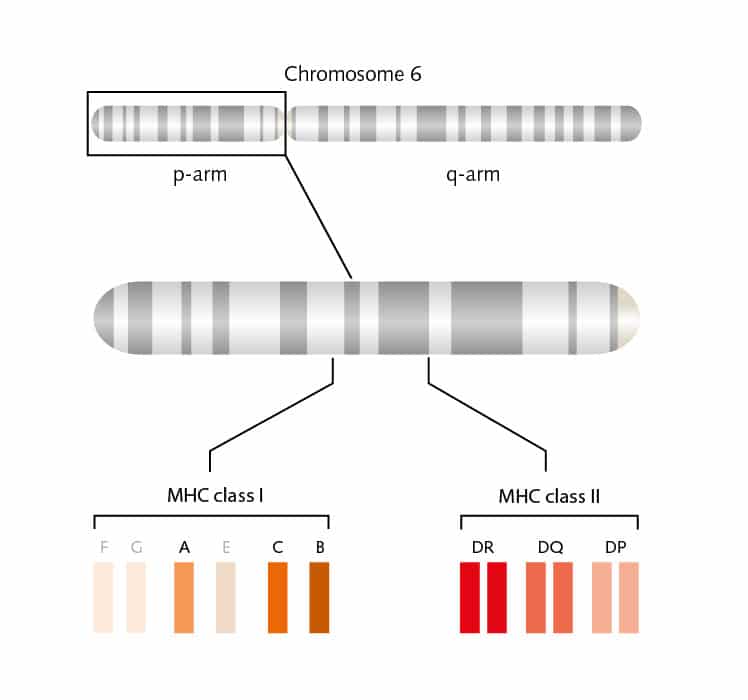
HLA restriction: finding and binding
More than 60 years ago, researchers observed that blood serum from one person reacted with white blood cells of another person. This led them to identify the first HLA molecule, HLA A-2 (Dausset, 1958). By 1968, so many HLA antigens had been identified that the HLA Nomenclature Committee was founded, with the aim of giving official names to new HLA molecules (Thorsby, 2009). In the 1970s, the Nobel-prize winners Rolf Zinkernagel and Peter Doherty showed that MHC enabled T cells to recognize antigens. However, researchers needed another decade to discover the rules for the interactions between HLA proteins and T cells, and how to explain the phenomenon of HLA restriction (Bjorkman et al., 1987; Rammensee et al., 1986). In fact, T cells can recognize peptides only if they are bound to HLA molecules. These peptide-HLA complexes interact only with HLA-matched T cells that have the corresponding receptors.
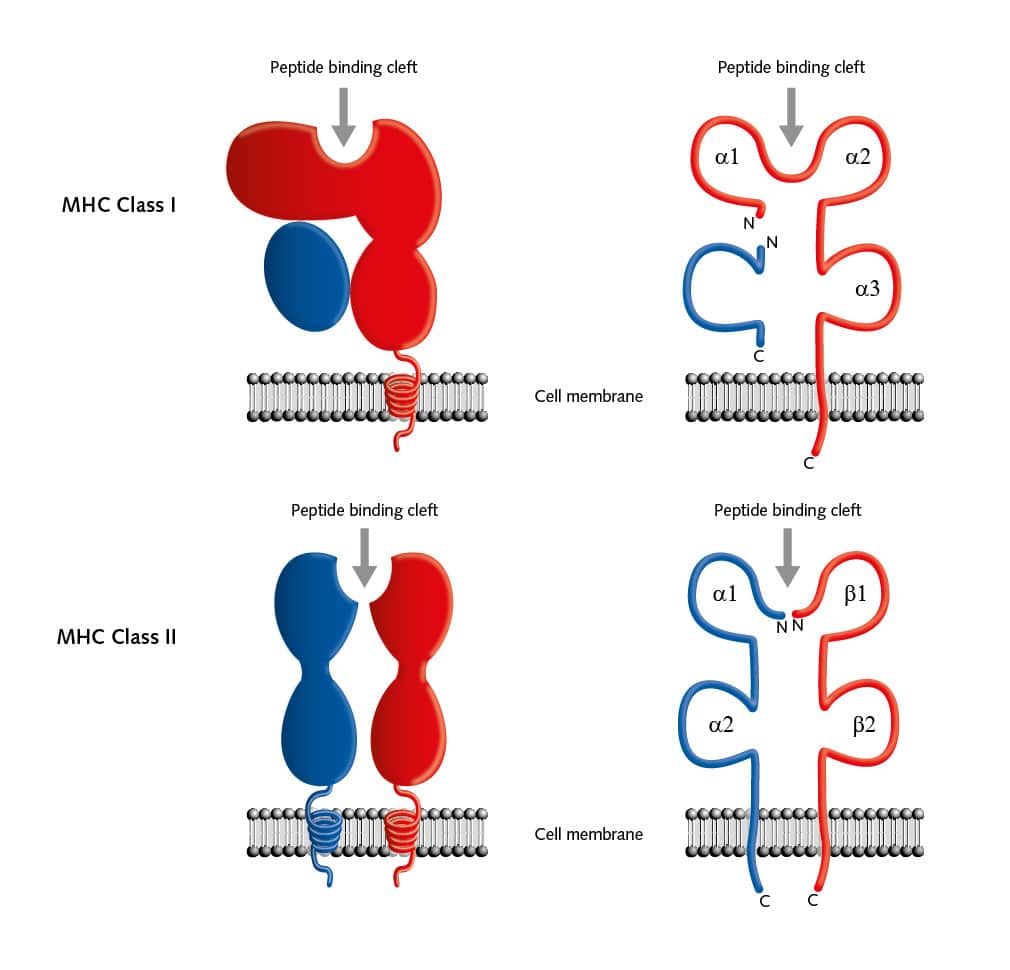
But there are many more pieces to this complicated puzzle. Two main classes of MHC molecules, MHC class I (HLA-A, HLA-B, and HLA-C) and MHC class II (HLA-DP, HLA-DR, and HLA-DQ) exist. Located between the genes coded for MHC class I and II proteins on chromosome 6, is a cluster of genes for MHC class III proteins. These proteins are not involved in antigen presentation, but mainly serve as signals for intercellular communication. MHC class I proteins are expressed on all cells of the body, except for red blood cells, and these proteins bind peptides from within the cells. MHC class II proteins are usually on the cell surface of antigen-presenting cells (APCs) and bind antigens that have been phagocytosed and processed from outside the body.
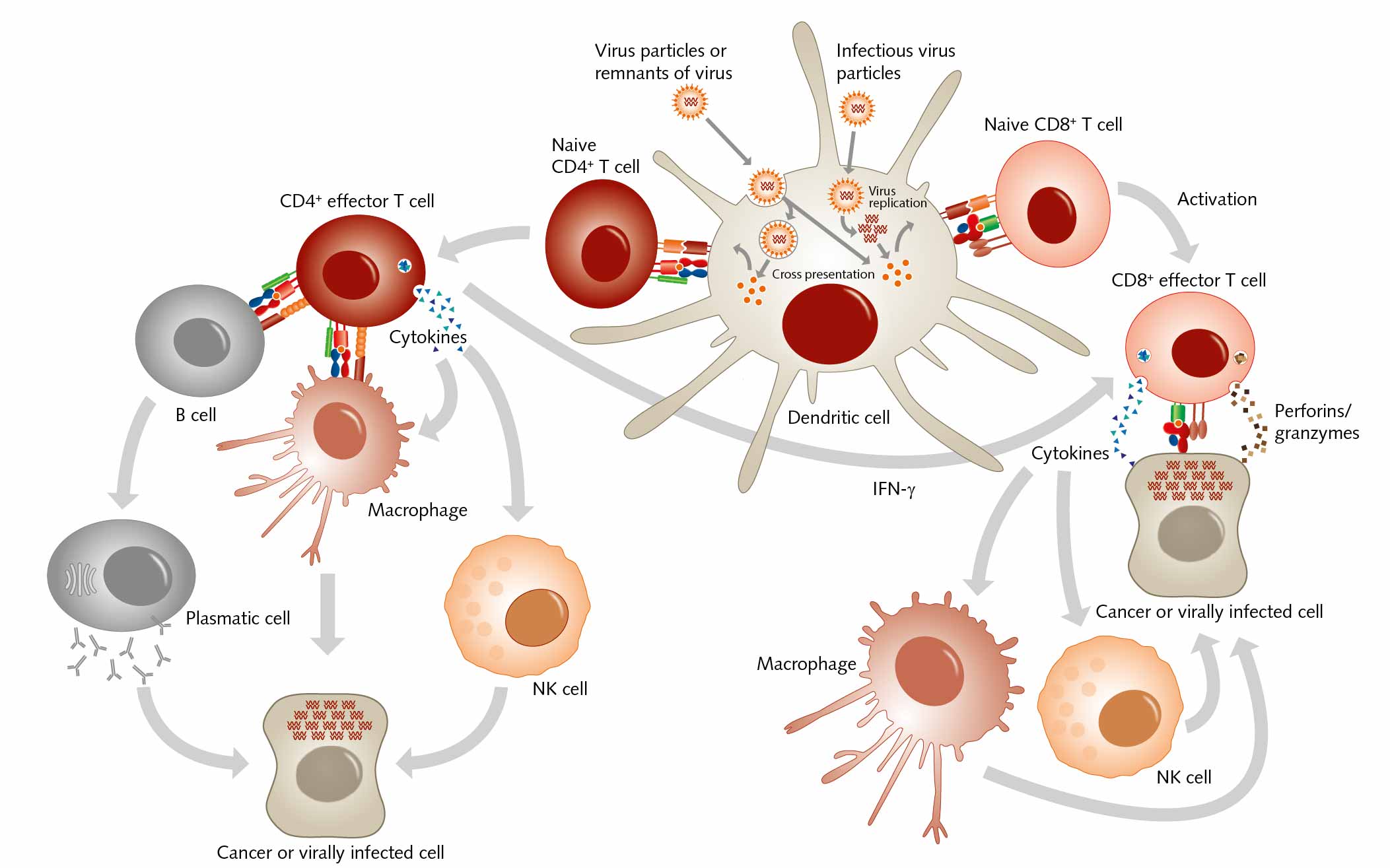
The structure of both MHC classes displays a groove where antigens, that is, processed peptides, can bind. T cells recognize the MHC-antigen complex and interact with it via the T cell receptor. Then APCs such as dendritic cells load proteins from extracellular sources, for example, loading infectious virus particles onto MHC I. When this MHC I/ antigen complex binds to the appropriate T cell receptor, this activity induces the differentiation of naive CD8+ T cells into CD8+ cytotoxic effector T lymphocytes. When these effector T cells are activated, they secrete lytic proteins that induce cell lysis or apoptosis of virus-infected cells. This action deprives the virus of the environment it needs to reproduce and survive. Effector T cells also produce cytokines, primarily TNF-α and IFN-γ, which activate other immune cells as macrophages or natural killer cells. And APCs can present peptides on MHC class II to CD4+ T helper cells, which activate other cells of the immune system such as B cells, macrophages, natural killer cells, and CD8+ T cells, for example by secreting cytokines.
HLA and diseases: activating autoimmune responses
Yet the immune response can backfire and having a specific HLA allele can hang like the sword of Damocles over a person. “It is true that HLA molecules are key for a functioning immune system. Yet, certain HLA alleles are associated with distinct pathological conditions,” remarks Schmidtke. Autoimmune diseases including rheumatoid arthritis, ankylosing spondylitis, type 1 diabetes, and Graves’ disease are related to certain HLA class I molecules. When cytotoxic T cells join in, they can spark the onset of the disease or worsen it. Variations in HLA class I genes can trigger autoimmunity, in particular, following bacterial or viral infections. Then the vicious circle becomes more vicious.
Microbial antigens, which are similar to self-antigens, can activate autoreactive T cells, which then cross-react with self-antigens and start attacking other organs in the body (Gough and Simmons, 2007). Many chronic and debilitating autoimmune disorders can’t be treated adequately, as no specific therapeutic options exist. Therapy then relies primarily on highly toxic drugs with severe side effects. “When we understand more about the specific immune mechanisms behind the development of autoimmune diseases, we could use immunotherapies more effectively. These support the body’s defense mechanisms without severe side effects,” explains Schmidtke.
Also, scientists suspect that specific HLA alleles make patients more susceptible to severe infections – or, conversely, protect them from such infections (Kenney et al., 2017). To elicit the desired immune response, both HLA/microbial antigens complexes and fitting T cells must exist. When either of these is missing, people tend to develop severe infections. On the other hand, patients with the right combination of HLA and T cells are often more resistant to infections. This is the case, for example, in malaria and HIV/AIDS, when specific HLA alleles protect the person from the infection (Mosaad, 2015).
HLA and cancer: how cancer cells escape recognition
The HLA profile can initiate the onset of cancer, or influence the response to chemotherapy, especially in virally-associated cancers (Little et al., 1999). Carcinogenic viruses activate genes that allow cancer cells to escape immune surveillance and proliferate without control. Possible immune escape mechanisms include changes in the structure and function of HLA, loss of expression of tumor antigens, and production of immunosuppressive cytokines. How well cancer patients respond to immunotherapy also depends on their HLA type, as patients with certain HLA class I molecules tend to have better outcomes. In a recent study, scientists observed the response of 369 melanoma patients to immune checkpoint inhibitors. Patients with the HLA-B44 type lived longer than those with the HLA-B62 (Chowell et al., 2017).
New therapeutic options: harnessing HLA for cancer immunotherapy

The body’s own peptides that are bound to HLA molecules on healthy cells usually don’t generate a T cell response. However, tumor cells contain additional tumor-specific peptides that aren’t present on normal cells. These tumor-associated antigens (TAA) can be encoded by mutated genes (neoantigens) or can be derived from proteins that are overexpressed in tumors. When presented by HLA molecules, TAA can activate cytotoxic T cells. These may be able to destroy the tumor cells before they proliferate or metastasize. The immune system’s ability to recognize TAA is where immunotherapy begins.
Stem-cell transplantation, a life-saving procedure for patients with hematological malignancies, found in bone marrow, for example, was one of the first attempts at immunotherapy. The success of a transplantation strongly depends on the HLA-compatibility between donor and recipient. If they are incompatible, this generates complex immune reactions. The body will reject the graft or cells will lose their function. To avoid rejection, as well as graft-versus-host-disease, HLA typing must be followed by matching of donor and recipient (Howard et al., 2016).
In cancer immunotherapy, neoantigens are very attractive targets. They are expressed only in malignant cells, and the T cell repertoire that recognizes them is not affected by central tolerance. Neoantigen-reactive T cells have been found in many tumors, and the number of neoantigens on cancer cells directly correlates to immunotherapy success (Karpanen and Olweus, 2017). However, sometimes autologous antigen-specific T cells don’t recognize neoantigens and they fail to stop cancer from spreading. This happens when T cells are inhibited or deleted by tolerance-inducing mechanisms.
Alternative approaches, such as using T cells from HLA-matched donors, can cure hematologic malignancies – because these cells recognize that polymorphic peptides in the donor and patient are different (Falkenburg, 2010). Adoptive immunotherapeutic approaches, as the transfer of CAR T cells, use genetically modified T cells from healthy (HLA-matched) individuals. The CAR T cells then respond to neoantigens that were ignored by the patient’s tumor-infiltrating T cells (Khalil et al., 2016). Even though neoantigens can be identified quickly, it takes time and resources to assess their immunogenicity. By finding tumor-specific biomarkers, along with greater understanding of each patient’s HLA type, researchers can slowly piece together the puzzle of possible therapies for each individual.
HLA-typed cells: an important control system to test new immunotherapies
Scientists are developing a range of strategies to defeat cancer. These strategies include using patients’ T cells, HLA-matched T cells, genetically modified T cells, or T cell receptors that recognize HLA-bound TAA and eliminate cancer cells.Before using T cells in patients, researchers must be sure that these cells will not attack healthy tissues and organs nor have any cytotoxic effects. To do this, they need to test T cells on HLA-matched primary cells from vital organs. “We offer HLA-typed cells from organs such as lung, heart, kidney, or the cardiovascular system, which allow scientists to test their therapies and detect eventual cross-reactions,” explains Lisa Schmidtke, Scientific Support Specialist at PromoCell. “HLA-typed cells provide an optimal control system to test cancer reactive T cells in cell culture. Animal tests don’t provide the same relevant evidence, as results cannot be directly transferred to the human system.”
Researchers can also use HLA-typed cells to develop new immunotherapies and to look for new antigens or biomarkers. These are convenient and save time because they needn’t to be screened. “HLA-typing of primary cells is a long process,” says Schmidtke. “Cells have to be first thawed and cultured before extracting the DNA. The genetic material is then sent to a lab for molecular typing. After a couple of weeks, the results are ready and if the HLA type of the tested cells does not match, a new donor has to be found and the process starts again.” Speeding up testing can also shorten the development time of new therapies for cancer patients.